GenEvo Curriculum
A computational systems biology curriculum for high school students to engage with cutting-edge ideas in modern biology
Project Overview
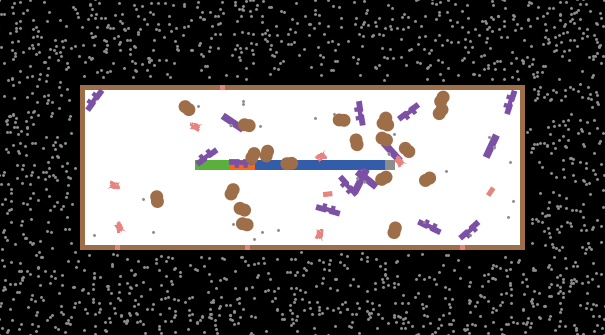
Fig1. A screen-capture of the Genetic Switch model
Did you know that bacteria are capable of making complex decisions? How do scientists use computers and engineering principles to change how bacteria act and understand their survival mechanisms? Computational modeling helps scientists, health care professionals, and others more deeply understand complex questions about natural and social phenomena.
This is a computational biology course that uses model-based-inquiry approach for learning. Students engineer genetic circuits using a computer simulation of a bacterial cell to develop deep understanding of cutting-edge ideas in genetics, evolution and complex systems thinking. This course fosters scientific thinking and computational thinking in the context of biological systems.
Essential Disciplinary Questions
- How do molecular interactions between genes and proteins result in regulating an organism’s response to the environmental changes through the feedback mechanism?
- How do populations evolve in nature under different environmental conditions?
- How do mechanisms of natural selection and genetic drift result in evolutionary changes at the short time scales (microevolution)?
Course Learning Outcomes (Based on Next Generation Science Standards):
Upon successful completion of this course, students will learn
Disciplinary Core Ideas:
- Genetic Regulation
- Feedback Mechanism
- Homeostasis
- Evolution - Natural Selection, Genetic Drift
Science Practices:
- Developing and Using Models
- Planning and Carrying Out Investigations
- Using Computational Thinking
- Constructing Explanations and Designing Solutions
- Engaging in Argument from Evidence
- Obtaining, Evaluating and Communicating Information
Crosscutting Concepts:
- Cause and Effects
- Systems and Systems Models
Download GenEvo Models
GenEvo Curriculum is included in the Models Library of NetLogo Version 6 and later. There are 5 models in this curriculum.
1. GenEvo 1 Genetic Switch
This model simulates a complex phenomenon in molecular biology: the “switching” (on and off) of genes depending on environmental conditions. Through molecular interactions between specific regulatory proteins and specific DNA sequences, each regulated gene is turn on or off in response to environmental stimuli. Download GenEvo 1 Genetic Switch.
2. GenEvo 2 Genetic Drift
This model is an example of genetic drift in a population of asexually reproducing bacteria E. coli. It starts with several types of E. coli, each with a different trait (which we represent with unique colors). The model shows that competing types of E. coli, each reproducing with equal likelihood on each turn, will ultimately converge on one type without any selection pressure forcing this convergence. The idea, explained in more detail in Dennett's Darwin's Dangerous Idea, is that trait drifts can occur without any particular purpose or 'selection pressure'. Download GenEvo 2 Genetic Drift.
3. GenEvo 3 Genetic Drift and Natural Selection
This model allows for the exploration and comparison of two different mechanisms of evolution: natural selection and genetic drift. It models evolution in a population of asexually reproducing bacteria. Download GenEvo 3 Genetic Drift and Natural Selection.
4. GenEvo 4 Competition
This is a population model of competition between cells in a population of asexually reproducing E. coli bacteria for resources in an environment. Users can experiment with various parameters to learn more about two different mechanisms of evolution: natural selection and genetic drift. Download GenEvo 4 Competition .
5. Genetic Switch - Synthetic Biology
This a multi-agent model of a genetic circuit in a bacterial cell and is an extension of the GenEvo 1 model. This model shows how biologists can use laboratory techniques to tweak certain aspects of a genetic circuit in order to affect the cell's behavior. Download GenEvo - Synthetic Biology.
Researchers
Sugat Dabholkar and Uri Wilensky
